Ancient enzymes reveal the DNA genesis
Recently found pre-enzymes give answers to the mystery of ancient amino acids assembling spontaneously to make up the first living cells
news | June 26, 2015

On June 18, 2015 the Journal of Biological Chemistry published a paper titled “Functional Class I and II Amino Acid Activating Enzymes Can Be Coded by Opposite Strands of the Same Gene” on the reconstruction of the Urzymes and protozymes which probably led to the appearance of modern life. We asked one of the authors of the research, Dr. Charles W. Carter Jr., to comment on this work.
The Study
Nature made at least three new types of inventions in assembling living cells from building blocks produced by prebiotic chemistry: catalysis synchronized the necessary chemical reactions; specificity put the building blocks together correctly; and heritable blueprints – genetic coding – furnished sufficient continuity for complexity to grow. The most dramatic of these inventions were all completed and probably overwritten before the first living cells appeared, making it difficult to recover relevant molecular fossils. Further, their daunting molecular diversity makes it hard to write a credible story line from fossils we do recover.
Our work chips away at all three inventions by focusing on one of the most difficult steps, chemically activating amino acids so that they can assemble spontaneously. That step is the slowest and most in need of being speeded up. How that happened is obscured by the wide range of amino acid physical properties like size, shape, and solubility in water versus oil. Each requires a different modern enzyme, called an aminoacyl-tRNA synthetase (aaRS), to make aminoacyl-tRNA. To make matters worse, aaRS come in two unrelated families; 10 of the 20 amino acids need a Class I aaRS, the other 10 a Class II aaRS. This landscape is thus littered with perplexing questions like these:
I. Why wasn’t one ancestor enough when they both do the same job?
II. How did the two types of ancestral synthetases avoid competition that might have eliminated the inferior Class?
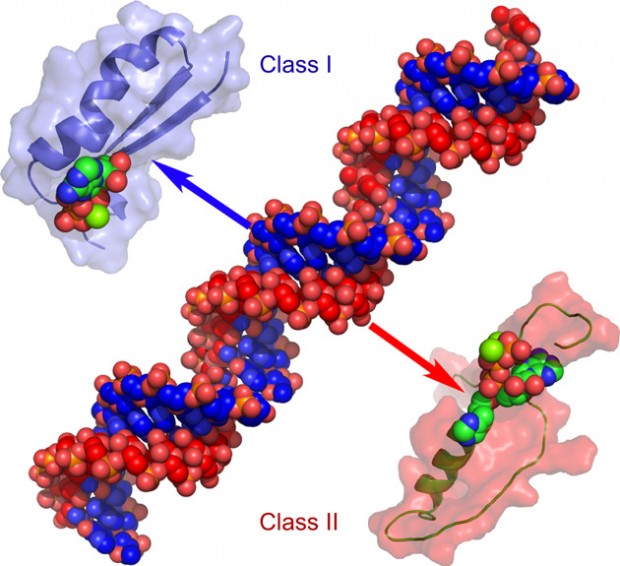
Computer model of urzymes and protozymes
Working by analogy with paleogenetics, we recovered successively smaller, more broadly conserved fossils by aligning the 3D structures of both superfamilies. We call them Urzymes (from Ur – ‘authentic’) and protozymes (from protos – ‘first’). We then devised a way to show experimentally that the blueprints for ancestral Class I and Class II protozymes are complementary versions of precisely the same information. The molecular machines made by interpreting the genetic blueprints and their complements are essentially ‘inside out’. Both work equally well in the test tube. The two families are so intimately connected that survival of one literally assured survival of the other.
This new unification of biology has many implications. Genome sequencing has produced the entire human blueprint. How many modern genes ultimately descended from one or the other strand of this ancestral, dual coding gene? My guess is roughly half. The fact that the protozymes are so active may also be relevant to human health: techniques we developed can potentially create compact forms of large genes that are too big to fit into the viruses used to deliver them for gene replacement therapy.
Backround

Much of my recent work has focused on deconstructing the two aaRS superfamilies in search of smaller, more broadly conserved modules to represent molecular ancestors. The work led first to the discovery that Urzymes derived from both Classes use only ~25% of the mass of modern aaRSs to produce 60% of their activity. Catalysis by aaRS protozymes now shows that proficiency – the logarithm of the rate acceleration – of a catalyst depends linearly on its mass.
Richard Wolfenden and I showed recently in the Proceedings of the National Academy of Sciences that the properties of the 20 amino acids are related closely both to protein folding and to synthetase-tRNA recognition elements. The key difference between amino acid substrates of Class I and Class II synthetases is that the former tend to form the insides, whereas the latter tend to form the surfaces of folded proteins, thereby rationalizing the need for two types of synthetases. Further, the tRNA acceptor stem recognition elements form a complete code for the sizes of the twenty amino acids; that code would therefore have been consistent with the requirements of early peptides to bind to RNA. The anticodon, which likely came along later, encodes amino acid polarity.
Future Directions

A central tenet of the evolutionary emergence of the genetic code is that it arose from a lower fidelity system. The unique information in the 46mer may be only 23 amino acids long as the coding DNA may be a palindrome. A more ambitious task now on the horizon is thus to show experimentally how something similar to the sense/antisense protozyme gene characterized in this paper could have arisen from a more random ensemble of simpler ancestors. Can we generate functional 23 residue ensembles according to a simpler code specifying a small number of different types of amino acids? Such questions were only theoretical before our work. Now they can be answered experimentally.
If you would like to contribute your own research, please contact us at [email protected]